- 4192 Devonwood Way, Ashburn, Virginia, 20148
- Helpline: +1 (703) 665-3747
Decision-Support Expert System to Assess Severe COVID-19
- Home
- Back to Journal
- Article Details
Abstract
Since the original outbreak of the disease there have been billions of people impacted, especially with severe COVID-19 cases. In fact, there are millions that died from COVID-19. However, there still remains to be seen a publically available chatbot for people, patients and care providers to determine the potential severity of a person’s COVID-19 infection and a more in-depth understanding of the human body system, co-morbidities and medical conditions that may become a severe case of COVID-19. Our preliminary investigation took this into consideration during a global pandemic and constructed a simple decision tree with binary classification that was stratified by age and body system, viral infection, co-morbidities, and any possible manifestations. We also reviewed several articles in the literature with special emphasis on manifestations. Moreover, we reviewed several chatbots currently utilized for COVID-19 to correlate any decision tree structures and frameworks that might be used for severe COVID-19 cases. The decision tree was constructed using the VisiRule© suite of tools to construct chatbots, decision tree frameworks and analytics. We managed to establish 212 nodes that were stratified from lung to heart conditions along body systems, medical conditions, co-morbidities and relevant manifestations that the literature indicated during the pandemic. This resulted in a possible 63,360 scenarios to end point of possible COVID-19 case. These scenarios provide a method to understanding the data needed to validate the decision tree and shows the complicated nature of severe cases of COVID-19. The decision tree does indicate that stratification of the viral infection with body system strengthened the framework to incorporate comorbidities and manifestations. However there are limitations. Using a decision tree does not provide for a circular reference of diagnosing COVID-19 based on different conditions, which makes the application less dynamic to a real-time diagnosis. Also, the decision-support is not based on data but information in published research during the pandemic, which we know some information rapidly changed in terms of how COVID-19 would impact us, especially for severe cases and hospitalizations during the major waves of the infections. Nevertheless, the VisiRule© application can provide modeling aspects as well as provide insight into the type of data required for a decision-support tool for severe COVID-19 assessments.
Keywords: chatbot, clinical decision support, co-morbidities, COVID-19, decision tree, manifestations, ontology-based stratification, prototype, and simulation
Introduction
The illness known as novel coronavirus disease 2019, or COVID-19 for short, presents a major and urgent threat to global health. COVID-19 is caused by a beta-coronavirus officially designated by the World Health Organization as SARS-CoV-2. Since the original outbreak of the disease in early December 2019 in Wuhan, China, COVID-19 has spread to 188 countries, with the number of cases recently estimated at ~50 million and the number of fatalities at ~1.2 million [1, 2, 3]. With that, approximately 3% of persons infected by the coronavirus have died [4]. In Canada, 30–40% of Canadians would have contracted COVID-19 and the proportion of the infected population whose illnesses are classed as severe is ~10% [5]. Moreover, there is increasing concern that severe cases may be increasing, despite the public-health efforts to contain the disease and inhibit its spread. An efficient expert system for the rapid assessment of severe COVID-19 cases would greatly benefit clinicians in dealing with this disease.
The sources and modes of transmission of COVID-19 remain unpredictable [1].
Nonetheless, scientific knowledge of the biology of the virus is well established. Coronaviruses are enveloped, single-stranded ribonucleic acid (RNA) viruses with surface spike proteins that penetrate the cellular cytoplasm of host organisms and transform cells for replication and shedding [6, 7]. COVID-19 is highly contagious, with a transmission capacity far greater than that of the previous SARS epidemic in 2003, and the virus reaches an extremely high abundance in infected people (up to a billion RNA copies/mL of sputum) [8]. Moreover, COVID-19 is stable on plastic and steel objects, viable virus remaining detectable for more than 72 hours [8]. The wide and rapid spread of COVID-19, together with its mortality rate, has occasioned a great deal of research on its physiological and health effects, including its manifestations in conjunction with existing morbidities. Much of this research is aimed at making accurate causal inferences with a view towards predictive modelling. Yet, the physiological interaction between COVID-19 and the human body is not yet well established. Research models need to be reframed or even re-invented based on an assessment of the problems and strengths of using a number of intersecting knowledge bases concerned with human pathophysiology against COVID-19.
Mild cases of COVID-19 usually have an early viral clearance, with 90% of mildly affected patients repeatedly testing negative on reverse transcriptase polymerase chain reaction (RT-PCR) by day 10 post-onset [9]. In contrast, all severe cases still tested positive at or beyond day 10 post-onset, with the median duration of viral shedding being 20 days (interquartile range: 17–24) in survivors, which is also the time when the affected individuals were most contagious [10]. In one study, the longest observed duration of viral shedding in survivors was 37 days [11], while other studies reported even longer durations [12]. Therefore, the determination of mildness or severity is an important aspect of diagnosis, because mild cases can quickly change into severe cases, especially if recovery is slow. Thus, the rapid assessment of COVID-19 cases as severe will aid clinicians in dealing with patients whose recoveries are delayed. Conversely, if a case is quickly diagnosed as mild, then the use of a ventilator can be ruled out; this is important because some studies have shown that intubation and mechanical ventilation can cause further chronic respiratory problems [13]. Since there is as yet no drug or vaccine capable of halting the COVID-19 pandemic, systems enabling rapid and reliable decision support are urgently needed. Such systems can be designed using data mining, machine learning, deep learning, and artificial- intelligence techniques as expert systems.
Current scientific research in the epidemiology of COVID-19, as well as public-health surveillance, relies on modelling COVID-19 infections to predict outbreaks. Recent studies have used medical images and radiology techniques to develop prognostic and diagnostic models for detecting COVID-19 using applied statistics and machine learning [9, 14]. Radiology is very important because it quickly detects COVID-19 as a respiratory disease at first onset but complications can occur, including cardiovascular diseases [9]. However, even though radiological diagnostic techniques offer a quick indication of COVID-19 severity, they do not have a high success rate in detecting severe cases within the important 10-day latency-to- replication phase, but there are “COVID-19 vulnerability index” related to hospital admissions [15, 16]. Modelling of COVID-19 suggests that severe cases can result in chronic respiratory and cardiovascular conditions lasting months and even longer, with recoveries remaining only partial with no treatment [9, 10, 17, 18, 19]. Thus, uncertainties remain, and patients’ responses vary greatly; therefore, quick and accurate detection of severe cases is important.
As long as no cure for COVID-19 exists, the capacity to predict severity becomes critical. One systematic review of +100 models of COVID-19 found that none were successful either in modelling its outbreak or in predicting recovery versus mortality [10]. Nevertheless, one recent study suggests that blood samples can be used to verify cases of severe COVID-19 without requiring x-rays or other tissue samples [18]. Epidemiologists and other health professionals are currently working together to model risks using applied Bayes-theorem with machine-learning techniques. This approach promises to produce models that can accurately assess the probability of severe cases within a given infected population, as well as complications of cardiovascular diseases [10, 19]. Patients who develop cardiovascular conditions can be treated by a inserting a pacemaker, but this is not common [20]. Additionally, steroids such as dexamethasone, prednisone, methylprednisolone, and hydrocortisone are used to reduce inflammation, while mechanical ventilators are used as a last resort [21]. Although updates on available treatments are frequent, gaps in the clinical guidelines for treating COVID-19 remain [10] and few clinical decision support tools have been validated [22].
Reframing the Problem of Decision Support of Severe COVID-19
In the current pandemic, clinicians need reliable information on human pathophysiological responses to COVID-19. Models commonly applied in prognosis and treatment include the simple nomogram (i.e., a diagram between variables such as age, gender, renal function, medication dosage, and body weight), decision trees, score systems, and online tools that provide statistics on in-hospital deaths, the need for prolonged mechanical ventilation, and a composite of poor outcomes [23]. However, there is little agreement as to which modelling techniques are useful in a hospital setting [10, 24]. One study of recovery in severe COVID-19 cases, based on testing of blood samples, showed that decision trees using Gini coefficients are more effective than models using support-vector machines, logistic regression, or random-forest classification [18]. Moreover, several studies indicate that decision trees can define complex outcomes of COVID-19, especially for severe in-hospital cases [25]. Thus, decision trees can be a valuable to incorporate knowledge structures that support the effective treatment of severe COVID-19.
Predictive models recently verified used specific parameters as predictors in the diagnosis and prognosis of COVID-19: age, body temperature, lymphocyte count, and data obtained through lung imaging [24]. In addition, flu-like symptoms and neutrophil count were frequently used as predictors in diagnosis, while comorbidities, gender, presence of creatinine reactive protein, and overall amount of creatinine were frequently used in prognosis [24]. Findings in regard to the biology and differentiation of lung inflammation suggest that excessive quantities of cytokines result in inner (microvascular) and outer (macro-vascular) heart problems and elevated risk of mortality [19]. Therefore, the main diagnostic-to-prognostic problem is the level of integration of COVID-19 with human biology and the variable degree of immune response. That is, some immune responses are beneficial in the body’s fight against COVID-19 while others are detrimental [26]. The diagnostic problem of ascertaining the probabilities of mortality and recovery can be simplified using binary classification (allowing for a variety of recovery versus death scenarios). The problem here is that there may be a variety of manifestations, each with its own specific physiological sequence, which can severely impact the body’s lung and/or heart [27]. Therefore, the design of a clinical decision-support tool to diagnose severe cases of COVID-19 would have to take into account these manifestations. It would have to incorporate human pathophysiology with COVID-19 and any remaining gaps in that understanding.
The knowledge of COVID-19 manifestations is complex, since a range of chronic conditions associated with lung, cardiovascular and gastro-intestinal diseases (refer to Table 1). One study showed that preceding coronavirus outbreaks, such as severe acute respiratory syndrome (SARS) and Middle East respiratory syndrome (MERS), were associated with a significant burden of cardiovascular comorbidities [28]. Furthermore, diagnostic workup during the SARS infection revealed detailed changes in electrocardiographic results, sub-clinical left ventricular diastolic impairment, and troponin elevation, all of which varied widely among patients [29]. Therefore, designing an expert system to support clinical decision-making needs to take account of the interconnectedness of the human body’s respiratory and cardiovascular systems in the context of severe COVID-19 infection. A decision tree can accomplish this by synthesizing knowledge structures in the architectural construction of its branches and leaves, which can be directly derived from data sets or modified to employ inductive and deductive reasoning.
Table 1: COVID-19 Manifestations.
|
Date |
Study |
Study Link |
Study Type |
Sample Size |
Manifestation |
Frequency of Symptoms |
|
2020-05-12 |
Manifestations and prognosis of gastrointestinal and liver involvement in patients with COVID-19: a systematic review and meta-analysis |
Systematic Review |
6686; 35 studies |
Gastrointestinal manifestations |
Diarrhea: 8%; Nausea and vomiting: 10%; Loss of appetite: 23%; Abdominal pain: 4% |
|
|
2020-04-29 |
Gender-Based Disparities in COVID-19 Patient Outcomes: A Propensity-matched Analysis |
Retrospective Observational Study |
13710 |
Gastrointestinal manifestations |
Abdominal and pelvic pain : 2.99%; Nausea and vomiting: 3.65%; |
|
|
2020-04-27 |
Prevalence and Impact of Myocardial Injury in Patients Hospitalized with COVID-19 Infection |
Retrospective Observational Study |
2736 |
Cardiac manifestations |
Tachycardia 23.6%; |
|
|
2020-04-23 |
Gastrointestinal symptoms as Covid- 19 onset in hospitalized Italian patients |
Prospective Observational Study |
411 |
Gastrointestinal manifestations |
GI symptoms: 10.2%; Nausea: 4.3% Vomit: 3.8%; Diarrhea: 3.6%; Abdominal pain: 1.2%; |
|
|
2020-04-20 |
Addressing the Question of Dermatologic Manifestations of SARS-CoV-2 Infection in the Lower Extremities: A Closer Look at the Available Data and its Implications. |
Retrospective Observational Study |
88 |
Dermatological manifestations |
Cutaneous manifestations: 20.4% |
|
|
2020-04-15 |
Beware: Gastrointestinal symptoms can be a manifestation of COVID-19 |
Systematic Review |
2800; 15 studies |
Gastrointestinal manifestations |
Diarrhea: 7.5%; Nausea: 4.5%; Anorexia: 4.4%; Abdominal pain: 0.5%; Belching/reflux: 0.3%; |
|
|
2020-04-09 |
Cardiac and arrhythmic complications in patients with COVID-19. |
Systematic Review |
953; 5 studies |
Cardiac manifestations |
Shock: 7–8.7%; Arrhythmia: 16.7%; Acute cardiac injury: 7.2%; Chest pain: 3.4%; ST†depression on ECG: 0.7%; Heart failure: 23%; Hypotension: 1%; Heart rate > 125 bpm: 1%; Ventricular fibrillation/ventricular tachycardia: 5.9% |
|
|
2020-04-06 |
In-hospital cardiac arrest outcomes among patients with COVID-19 pneumonia in Wuhan, China |
Retrospective Observational Study |
761 |
Cardiac manifestations |
Cardiac arrest: 19.8% |
|
|
2020-04-04 |
High Prevalence of Concurrent Gastrointestinal Manifestations in Patients with SARS- CoV-2: Early Experience from California. |
Retrospective Observational Study |
116 |
Gastrointestinal manifestations |
GI symptoms: 31.9%; Nausea/vomiting: 10.3%; Diarrhea: 10.3%; Nausea/vomiting and Diarrhea: 4.3%; Abdominal pain: 8.8%; Loss of appetite: 25.3% |
|
|
2020-04-04 |
High Prevalence of Concurrent Gastrointestinal Manifestations in Patients with SARS-CoV-2: Early Experience from California. |
Retrospective Observational Study |
65 |
Hepatic manifestations |
Abnormal liver function test: 40% |
|
|
2020-04-03 |
Clinical Features of COVID-19-Related Liver Damage |
Retrospective Observational Study |
148 |
Hepatic manifestations |
Abnormal liver function tests: 37.2%; Elevated AST: 21.6%; Elevated ALT: 18.2%; Elevated GGT: 17.6%; Elevated ALP: 4.1%; Elevated total bilirubin: 6.1% |
|
|
2020-03-31 |
Review article: gastrointestinal features in COVID-19 and the possibility of faecal transmission. |
Retrospective Observational Study |
2,023 |
Gastrointestinal manifestations |
Anorexia: 39.9%- 50.2%; Diarrhoea: 2%-49.5%; Vomiting in adults: 3.6%-15.9%; Vomiting in children: 6.5%-.7%; Gastrointestinal bleeding: 4%-13.7%; Abdominal pain: 2.2%-6.0% |
|
|
2020-03-31 |
Characteristics of Ocular Findings of Patients With Coronavirus Disease 2019 (COVID-19) in Hubei Province, China. |
Retrospective Observational Study |
38 |
Ocular manifestations |
Ocular abnormalities: 31.6%; |
|
|
2020-03-30 |
Suspected myocardial injury in patients with COVID-19: Evidence from front-line clinical observation in Wuhan, China |
Retrospective Observational Study |
112 |
Cardiac manifestations |
Possible myocarditis: 12.5%; |
|
|
2020-03-27 |
Cardiovascular Implications of Fatal Outcomes of Patients With Coronavirus Disease 2019 (COVID-19) |
https://jamanetwork.com/ |
Retrospective Observational Study |
187 |
Cardiac manifestations |
Myocardial injury: 27.8% |
|
2020-03-27 |
Predicting COVID- 19 malignant progression with AI techniques |
Retrospective Observational Study |
133 |
Gastrointestinal manifestations |
GI symptoms: 24.8% |
|
|
2020-03-26 |
Gastrointestinal Manifestations of SARS-CoV-2 Infection and Virus Load in Fecal Samples from the Hong Kong Cohort and Systematic Review and Meta- analysis. |
Retrospective Observational Study |
59 |
Gastrointestinal manifestations |
GI symptoms: 25.4%; Vomiting: 1.7%; Diarrhea: 22%; Abdominal pain/discomfort: 11.9% |
|
|
2020-03-26 |
Clinical characteristics of 113 deceased patients with coronavirus disease 2019: retrospective study |
Retrospective Observational Study |
113 |
Neurological manifestations |
Hypoxic encephalopathy: 20% |
|
|
2020-03-26 |
Clinical characteristics of 113 deceased patients with coronavirus disease 2019: retrospective study |
Retrospective Observational Study |
113 |
Cardiac manifestations |
Acute cardiac injury: 77%; 49% |
|
|
2020-03-26 |
COVID-19 clinical characteristics, and sex-specific risk of mortality: Systematic Review and Meta- analysis |
Systemic Review |
4789; 32 studies |
Gastrointestinal manifestations |
Diarrhea: 5.7%; Nausea/Vomiting: 5.0 % |
|
|
2020-03-23 |
COVID-19 Myocarditis and Severity Factors: An Adult Cohort Study |
Retrospective Observational Study |
84 |
Cardiac manifestations |
Abnormal ECG and serum myocardial level: 15.48%; Myocarditis: 4.8% |
|
|
2020-03-20 |
Don’t Overlook Digestive Symptoms in Patients With 2019 Novel Coronavirus Disease (COVID-19) |
Retrospective Observational Study |
1141 |
Gastrointestinal manifestations |
Only GI symptoms: 16%; Nausea: 73%; Vomiting: 65%; Abdominal pain: 25%; Diarrhea: 37%; Loss of appetite: 98%; Both nausea and vomiting: 20%; Both abdominal pain and diarrhea: 9%; All GI symptoms combined: 7% |
|
|
2020-03-19 |
Characteristics and Outcomes of 21 Critically Ill Patients With COVID-19 in Washington State |
Retrospective Observational Study |
21 |
Cardiac manifestations |
Cardiomyopathy: 33.3% |
|
|
2020-03-16 |
Clinical characteristics of refractory COVID-19 pneumonia in Wuhan, China. |
Retrospective Observational Study |
155 |
Gastrointestinal manifestations |
Anorexia: 31.7%; Diarrhea: 4.5% |
|
|
2020-03-16 |
Ocular manifestations and clinical characteristics of 534 cases of COVID-19 in China: A cross- sectional study |
https://www.medrxiv.org/content/ |
Cross Sectional Study |
534 |
Ocular manifestations |
Conjunctival congestion: 4.68%; Dry eye: 20.97%; Blurred vision: 12.73%; Foreign body sensation: 11.80% |
|
2020-03-12 |
Association of Cardiovascular Manifestations with In-hospital Outcomes in Patients with COVID-19: A Hospital Staff Data |
Retrospective Observational Study |
41 |
Cardiac manifestations |
With cardiovascular manifestations: 58.5%; Without cardiovascular manifestations: 41.5% |
|
|
2020-03-11 |
Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study |
Retrospective Observational Study |
191 |
Cardiac manifestations |
Heart failure: 23%; Acute cardiac injury: 17% |
|
|
2020-03-08 |
Acute Myocardial Injury of Patients with Coronavirus Disease 2019 |
Retrospective Observational Study |
53 |
Cardiac manifestations |
Cardiac symptoms: 79.25%; Tachycardia: 28.3%; Electrocardiography abnormalities: 20.8%; Diastolic dysfunction: 37.7%; Elevated myocardial enzymes: 56.6%; AMI: 11.3% |
|
|
2020-03-06 |
Clinical features and outcomes of 221 patients with COVID-19 in Wuhan, China |
Retrospective Observational Study |
221 |
Gastrointestinal manifestations |
Anorexia: 36.2%; Diarrhea: 11.3%; Abdominal pain: 2.3% |
|
|
2020-03-06 |
Clinical features and outcomes of 221 patients with COVID-19 in Wuhan, China |
https://www.medrxiv.org/content/ |
Retrospective Observational Study |
221 |
Cardiac manifestations |
Arrhythmia: 10.9%; Acute cardiac injury: 7.7% |
|
2020-03-05 |
Clinical Characteristics of COVID-19 Patients With Digestive Symptoms in Hubei, China |
Cross Sectional Study |
204 |
Gastrointestinal manifestations |
GI symptoms: 18.6%; Diarrhea: 17%; Vomiting: 3.88%; Abdominal pain: 1.94% |
|
|
2020-02-29 |
Clinical Characteristics of Imported Cases of COVID-19 in Jiangsu Province: A Multicenter Descriptive Study |
Retrospective Observational Study |
80 |
Hepatic manifestations |
Increased ALT: 3.75%; Increased AST: 3.75% |
|
|
2020-02-25 |
Neurological Manifestations of Hospitalized Patients with COVID-19 in Wuhan, China: a retrospective case series study |
Retrospective Observational Study |
214 |
Neurological manifestations |
Neurological symptoms: 36.4%; CNS symptoms: 24.8%; PNS symptoms: 8.9%; skeletal muscle symptoms: 10.7% |
|
|
2020-02-21 |
Clinical course and outcomes of critically ill patients with SARS-CoV-2 pneumonia in Wuhan, China: a single-centered, retrospective, observational study |
https://www.thelancet.com/pdfs/ |
Retrospective Observational Study |
52 |
Gastrointestinal manifestations |
Vomiting: 4% |
|
2020-02-21 |
Clinical course and outcomes of critically ill patients with SARS-CoV-2 pneumonia in Wuhan, China: a single-centered, retrospective, observational study |
https://www.thelancet.com/pdfs/ |
Retrospective Observational Study |
52 |
Neurological manifestations |
Headache: 6% |
|
2020-02-19 |
Clinical findings in a group of patients infected with the 2019 novel coronavirus (SARS- Cov-2) outside of Wuhan, China: retrospective case series |
Retrospective Observational Study |
62 |
Neurological manifestations |
Headache: 34% |
|
|
2020-02-15 |
Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China |
Prospective Observational Study |
41 |
Cardiac manifestations |
Acute cardiac injury: 12% |
|
|
2020-02-07 |
Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus Infected Pneumonia in Wuhan, China |
Retrospective Observational Study |
138 |
Gastrointestinal manifestations |
Anorexia: 39.9%; Diarrhea: 10.1%; Nausea: 10.1%; Vomiting: 3.6%; Abdominal pain: 2.2% |
|
|
2020-02-07 |
Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus Infected Pneumonia in Wuhan, China |
Retrospective Observational Study |
138 |
Cardiac manifestations |
Acute cardiac injury: 7.2%; arrhythmia: 16.7% |
|
|
2020-01-30 |
Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study |
Retrospective Observational Study |
99 |
Neurological manifestations |
Confusion: 9%; Headache: 8% |
|
|
2020-01-30 |
Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study |
Retrospective Observational Study |
99 |
Cardiac manifestations |
Chest pain: 2% |
|
|
2020-01-30 |
Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study |
Retrospective Observational Study |
99 |
Gastrointestinal manifestations |
Diarrhea: 2%; Nausea and vomiting: 1% |
A Binary Decision-Tree to Assess Severe COVID-19
It is known that a decision tree can incorporate medical knowledge [30], including human physiological responses to influenza and other diseases [25]. Human immune responses to COVID-19 can reveal normal increase in T-cells due to inflammation; however, that then leads to a cytokine storm that increases the risk of mortality rather than reducing it. This makes construction of the diagnostic decision branches in the knowledge tree difficult, perhaps requiring the introduction of additional calculations, like Gini coefficient thresholds [28]. Also, the knowledge tree needs to include a transition from diagnosis of chronic lung to heart conditions, both microvascular (tissues and vessels) and macro-vascular (muscles and valves). A few clinical guidelines can be used in constructing the framework of decision-support chatbots that models the transition from mild to severe COVID-19 cases (Table 2). However, the stratification of interactions between COVID-19 and chronic lung conditions and heart disease is not well known [31].
Table 2: Different Chatbots for COVID-19 (environmental scan).
|
Author |
Ages |
Study Design and Description |
Setting |
Inference Method |
Intervention Effectiveness |
Guidelines Used |
Prescriptions |
|
Apple and Siri |
all |
Apple and Siri to Help people who ask if they have the coronavirus. |
Anywhere |
Uses data from John Hopkins University, rules based |
Only symptom based |
No clinical guidelines |
No |
|
Intermountain Healthcare |
all |
COVID-19 symptom checker |
Research, primary care, and acute care in Utah at Intermountain Healthcare |
Uses data from acute care settings, rules based |
Effective to separate mild and severe cases based on symptoms |
Simple clinical guidelines |
Unknown |
|
Google Dialogflow and Google Assistant |
all |
Chatbot designed with extensive prompts to entities and intents |
Anywhere |
Prototype, fact checker and QandA for COVID-19 |
Answers a variety of questions including data on COVID-19 and symptoms |
No clinical guidelines |
No |
|
COVIDradar |
all |
Chatbot and app to track health status |
Anywhere within UK and NHS |
Prototype, fact checker and QandA for COVID-19 |
Effective to separate mild and severe cases based on symptoms and daily updates |
Simple clinical guidelines |
Unknown |
|
Facebook Messenger with WHO |
all |
COVID-19 fact and symptom checker |
Anywhere |
Prototype, fact checker and QandA for COVID-19 |
Answers a variety of questions including data on COVID-19 and symptoms |
WHO guidelines |
No |
|
BC CDC |
all |
CDSS generated care suggestions based on agreed guidelines. These include what to do if negative tested for COVID-19 |
Research, hospital-based academic groups, possibly includes clinicians |
Prototype, fact checker and QandA for COVID-19 |
Answers a variety of questions including data on COVID-19 and symptoms |
Simple clinical guidelines |
No |
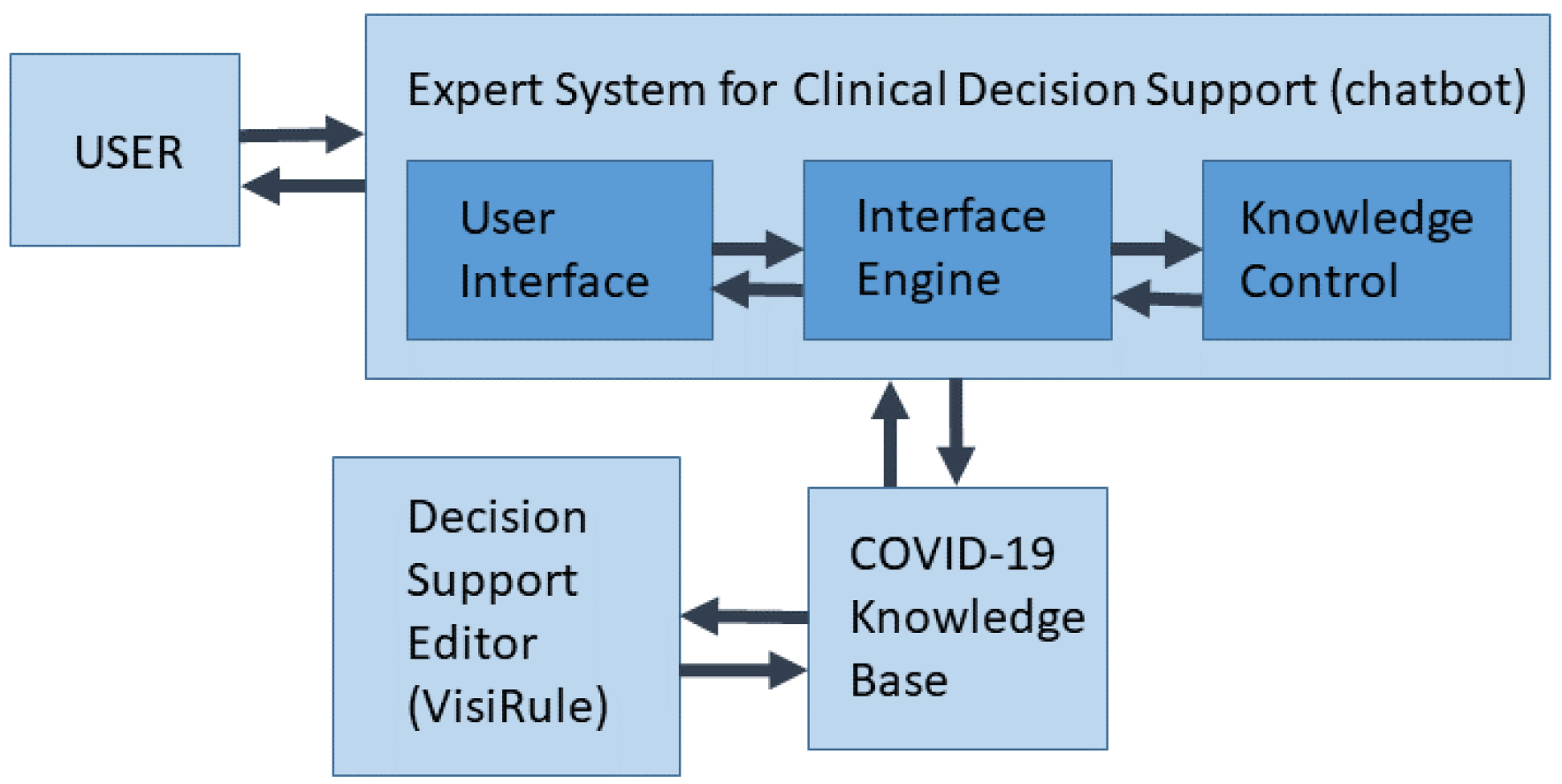
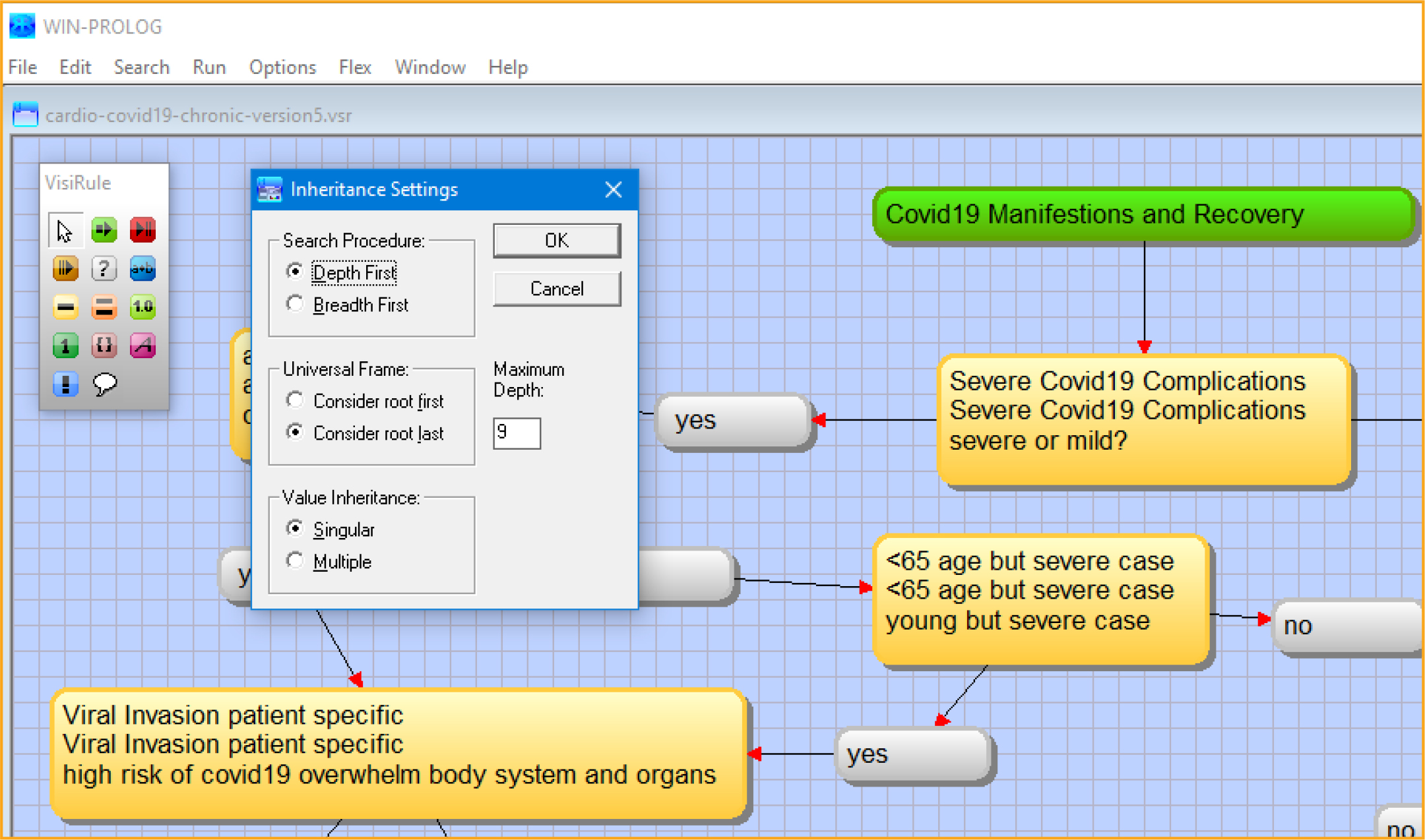
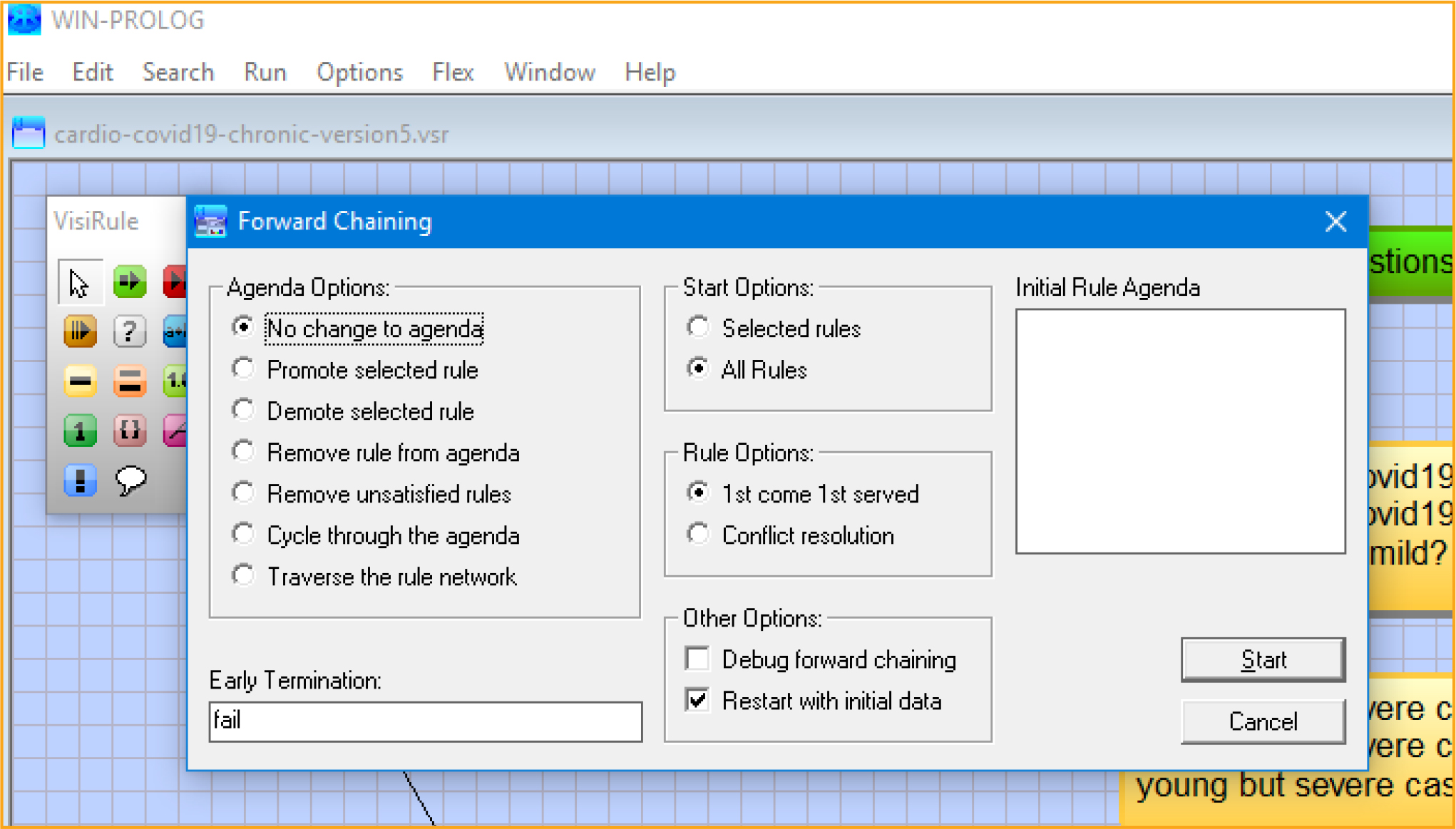
After reviewing expert systems, and other modelling technologies that use decision trees, we decided to use the infrastructure of a chatbot tool [32]. This tool can also use Predictive Model Markup Language (PMML) schematic formats to create a decision tree or it can also be used to design decision trees manually. For instance, PMML can be utilized in tools like KNIME© and RapidMiner© to automate the formation of decision tree; in such procedures, datasets and trained data can be employed iteratively. Moreover, it has been applied to binary classification in a variety of clinical applications (e.g., wound-care management) [32]. Fortunately, VisiRule© has knowledge engineering, interface and control/editor tools (Figure 1) with embedded inductive techniques with inheritance settings, set as singular “Depth First” with maximum of 9 from root node (Figure 2) with forward chaining settings (Figure 3). Induction (and probably deduction too) can be used in the diagnosis of coronavirus disease, an example being the use of the c4.5 algorithm in its sequence of decision points with decision endpoints [33]. In fact, the first approach used KNIME© to develop a decision tree based on existing COVID-19 datasets. The website Kaggle.com provides datasets on COVID-19 cases that then was used in the decision- tree framework in KNIME©, auto-generating a PMML schema. However, the COVID-19 datasets were not easily formed to binary classifications, specifically in terms of recovery, short or long recover with onset of chronic conditions, and even risk of mortality. This was confirmed by running Bayes modeling of the dataset in the Microsoft Azure Machine Learning Studio©, which did not produce accurate results. Accordingly, we constructed the decision tree to account for health outcomes, stratified by comorbidities in the transition from lung to heart, while establishing binary classifications to manual calculated Bayes probabilities for each leaf endpoint (Table 3).
Table 3: Risk factors for severe cases of COVID-19.
|
Risk Status |
Relative Risk (%) |
|
High Risk |
|
|
Personal history of multiple comorbidities including hypertension |
35 |
|
Previous lung cancer or chronic pneumonia |
17 |
|
Immune deficiencies |
5 to 15 |
|
Personal history of cardiovascular diseases |
9 to 10 |
|
History of heart issues like arrhythmia |
8 |
|
Immunosuppression |
6 to 8 |
|
Moderate Risk |
|
|
Asthma and Chronic Pulmonary Obstructive Disease (CPOD) |
4.9 to 7.3 |
|
Slight Inflammation |
3.0 to 5.4 |
|
Issues with coronary artery and micro-vessels |
5.4 |
|
Intermediate Risk |
|
|
Asthma but no chronic conditions but could have pneumonia or bronchitis |
3.8 |
|
Asthma, hypertension and slight inflammation |
3.0 |
|
Age < 65 years with micro-vessels and coronary artery issues |
2.2 |
|
Low Risk |
|
|
Age <65 years |
0.5 |
|
Normal level of COVID-19 replication for 10 days and test negative to covid- 19 |
0 |
Figure 1: Expert system of COVID-19 decision support web-based (chatbot) tool.
Figure 2: VisiRule inheritance settings, set as singular “Depth First” with maximum of 9 from root node. Screenshot from VisiRule 7.021.
Figure 3: VisiRule forward chaining settings. Screenshot from VisiRule 7.021.
We also tested Google’s Dialogflow application, which is essentially a decision-support tool in the form of a chatbot. This chatbot contains intents and entities and a knowledge base; its construction enables easy integration into web-based applications [34]. However, Dialogflow does not allow for binary classification; all it can do is enable a series of prompts in a specific sequence of COVID-19 queries. This makes the modeling of severe cases of COVID-19 extremely difficult in Dialogflow with absolutely no ability to incorporate Bayes theorem. In short, Dialogflow cannot be utilized. Another tool we considered, ZenChat©, proved similar to Dialogflow with no apparent capacity to generate decision trees using binary classification [35].
With the VisiRule© construct (via upload .vsr file to a website), prototypes were developed in which a report is generated after each iteration of a user selecting “yes” or “no” in a set of questions linked to a patient’s health and biological responses. The maximum run were about 10–12 leaves to an endpoint indicating high risk of mortality, low risk of mortality with risk of morbidity, or prolonged recovery from COVID-19 [36]. The tool allows the user to trace the response in the decision tree. Results are also obtainable in other forms, such as an HTML application that generates a list of results and a chatbot. Upon testing, it became clear that the decision tree used biological and physiological knowledge in deductive reasoning, with some inductive reasoning in the knowledge acquisition. However, the decision tree did not rebalance itself, and rebalancing will require additional datasets to allow for reordering the sequence of prompts, reducing or increasing the number of leaves leading to a decision point, and resampling data [30, 37, 38].
A script was run to generate “truth table” based on the number of 212 nodes of binary classification “Yes” in the VisiRule© application. The generated table consisted of 990 rows by 64 columns or 990 × 64 = 63,360 possible combinations to potentially contract COVID-19.
Inductive or inferential reasoning is the process of moving from concrete examples to general models in order to learn how to classify objects by analyzing a set of instances (e.g., cases of illness that have already been resolved) whose classes are known [37]. To improve the tool’s inductive power, it was decided to include asthma, pneumonia and chronic obstructive pulmonary disease (COPD) in the upper stratification of the decision tree, because these lung conditions can exacerbate cases of COVID-19 [13]. Furthermore, studies have linked acute complications of COVID-19 with an elevated long-term risk of cardiovascular disease [39]. This finding was incorporated into the decision-support tool after respiratory, as well as hyperlipidaemia responses, but gastro-intestinal responses were not included.
Objectives of Decision Support
In this study, the main objective of the decision-support tool was to diagnose severe cases of COVID-19 based on knowledge using VisiRule© control/editor tools (Figure 1). Decision support in both diagnosis and prognosis takes account of a variety of signs and symptoms of COVID-19, on the one hand, and of the ability of human biological responses to combat the virus, on the other (Table 1). In doing so, the clinical decision-support tool could help general practitioners, specialists, nurses, and physicians diagnose severe cases of COVID-19. The early detection even of mild cases of COVID-19 is important because it facilitates public-health measures that can lower the overall prevalence of severe cases [31].
The main objective of the decision-support tool, outlined above, can be broken down into a number of sub-objectives, as follows:
- Minimize severity of COVID-19 through early detection via binary classification;
- Assess severity of cases with the extent of respiratory and cardiovascular involvement;
- Reduce inaccuracies in the diagnosis of COVID-19, including both false negatives and false positives calculated by Bayes-theorem; and,
- Assess the risk of prolonged recovery, morbidity, and mortality.
In the process of achieving these sub-objectives, a clear set of clinical guidelines for dealing with COVID-19 will be achieved. Since early detection of COVID-19 is problematic, the percentage of mild and asymptomatic cases being high, several pathophysiological responses to the disease need to be fully documented in order for risk of severity to be adopted [10, 24].
Assessing severity, for instance, requires a high level of diagnostic accuracy, together with monitoring and periodic reassessment [19, 31], all of which are more easily accomplished with the use of a decision-tree tool such as the one described herein.
Strengths of Expert System
An effective clinical decision support (CDS) ensure that best practices are followed in the diagnosis and treatment of COVID-19 (refer to Tables 4–6). Among the benefits provided by a CDS are the following:
A CDS incorporates health outcomes of severe COVID-19 cases;
- A CDS takes into account of respiratory and cardiovascular symptoms related to severe COVID-19, (refer to Figures 4–6);
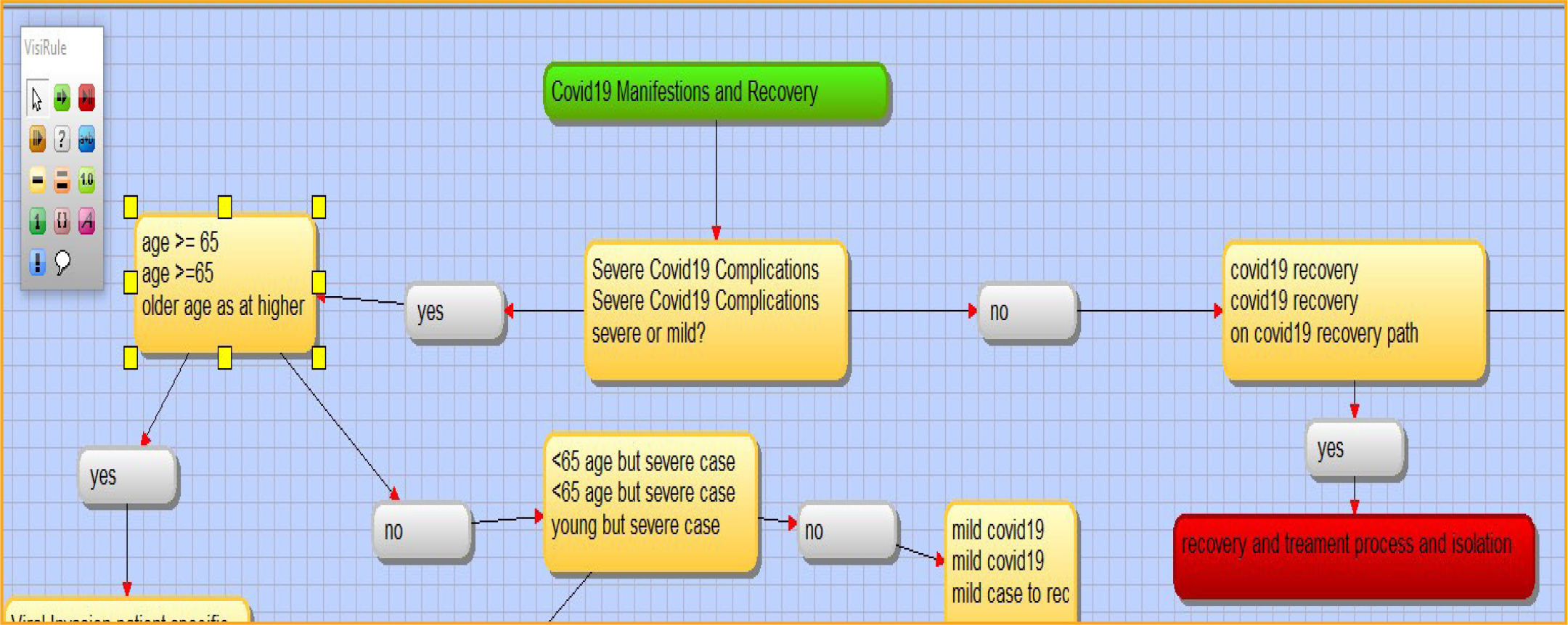
- Mild COVID-19 cases are initially separated from severe COVID-19 (Figure 7);
- The CDS can correlate COVID-19 infection with other indicators like obesity, diabetes, blood type, age, heart and lung complications and illnesses;
- The CDS can cover a wide gamut of human phytopathology issues relevant to severe COVID-19;
- A CDS uses binary classification in a decision-tree framework to predict risk of mortality;
- The tree map framework with the decision nodes (212) showed a large robust number of scenarios of 63,360 scenarios in the “Yes” binary classification to potentially contract COVID-19; and,
- The diagnostic and risk-stratification used in a CDS can be updated as knowledge of health outcomes and treatments is validated.
Table 4: A stratified approach to detecting severe COVID-19 cases.
|
1 |
Viral Invasion |
Suggestive of the level of the COVID-19 virus invading cytoplasm of cells and ACE2 |
|
2 |
Age and Body Systems |
Suggestive of higher risk of >= 65 years old with respiratory and circulatory systems impacted |
|
3 |
Comorbidities |
Suggestive of underlying chronic diseases |
|
4 |
Manifestations |
Suggestive of the level of viral infection and the human body’s responses |
Table 5: Seven-point checklist for severe COVID-19.
|
Type of conditions |
Parameter |
Interpretation |
|
Major conditions |
Change in lung |
One or more major signs: refer for expeditious increases or decreases in inflammation, T-cells and cytokines; additional presence of one or more minor signs increases the possibility of congestive heart failure. |
|
Change in heart |
||
|
Change in circulatory system |
||
|
Minor conditions |
Age < 65 years old |
Three or four minor signs without a major sign. |
|
Viral Invasion is in shedding phase in 10 days |
||
|
No Inflammation |
||
|
No pneumonia |
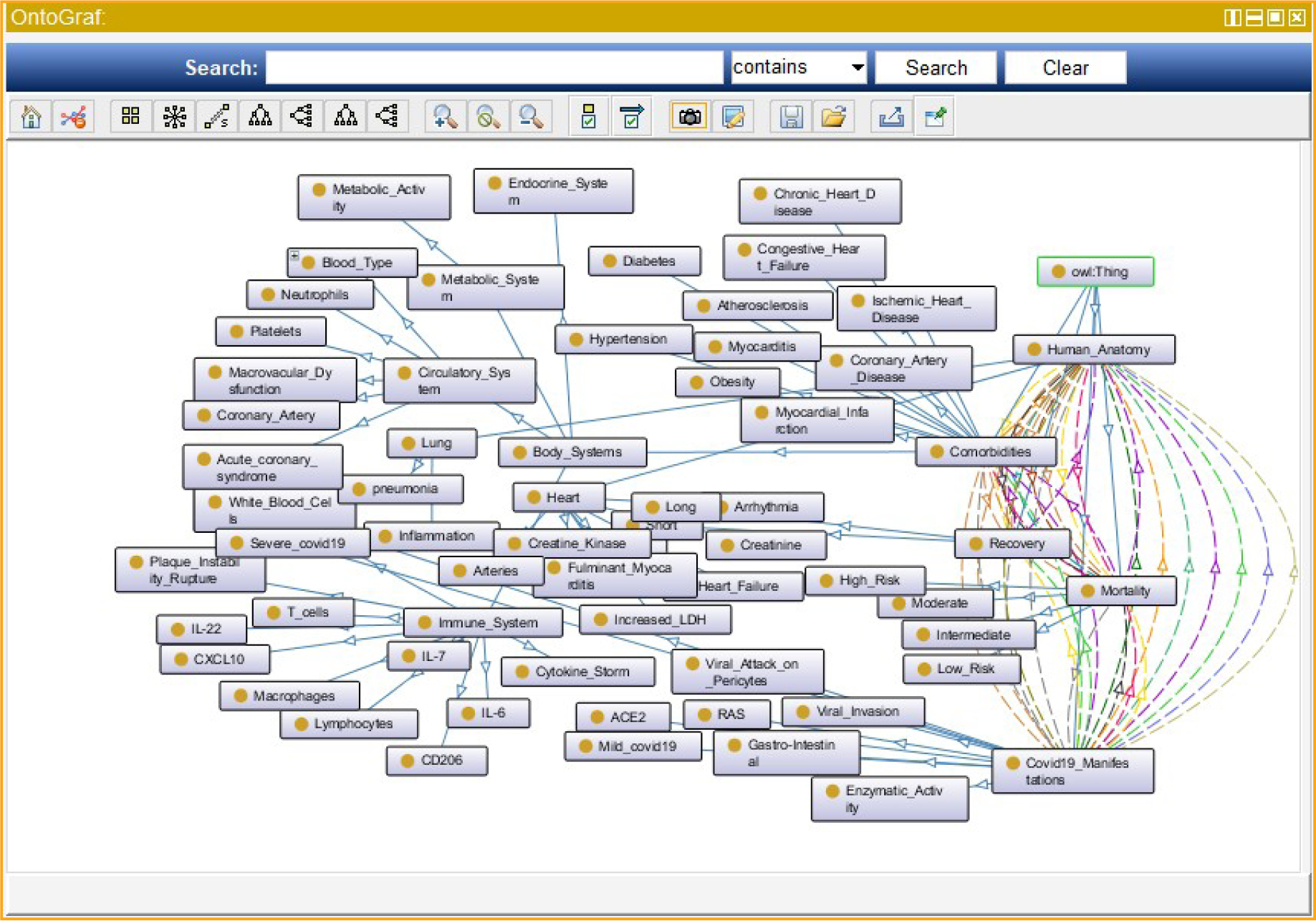
Table 6: List of Parameters of decision support (expert system) developed using Protégé 5.5.0 application.
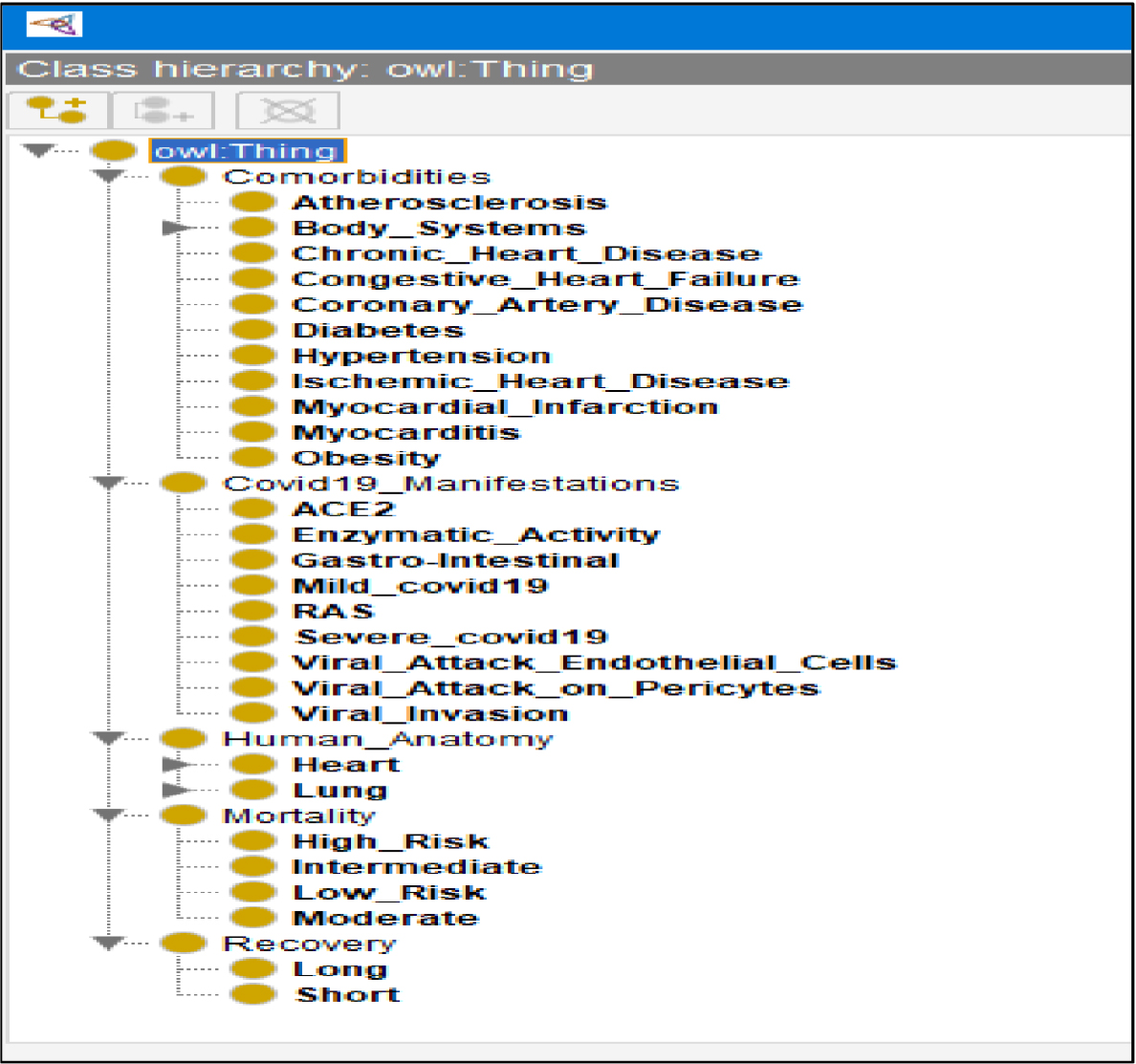
Figure 4: Ontologies of the COVID-19 in the decision tree of expert support toll using Protégé 5.5.0 application
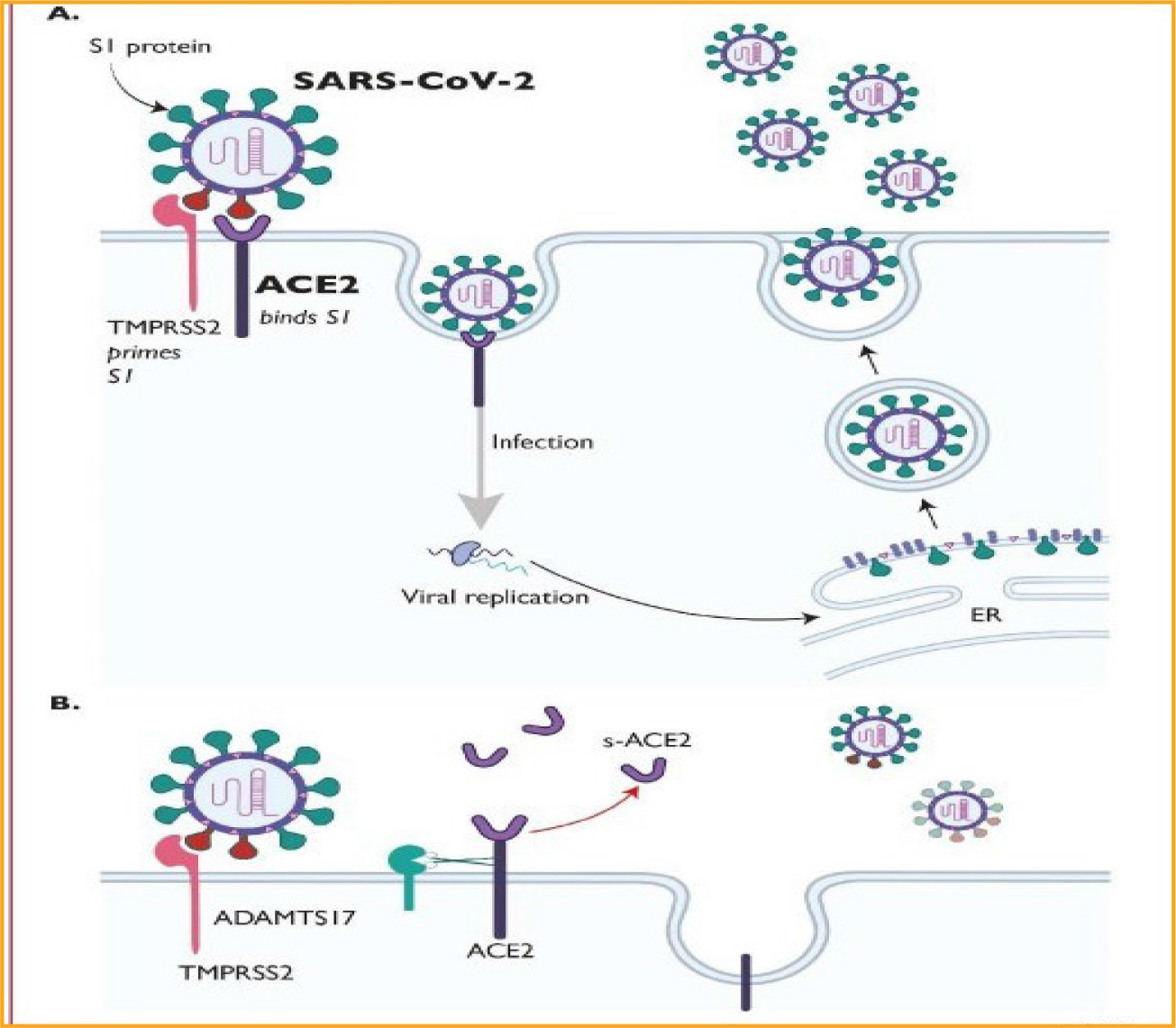
Figure 5: COVID-19 life cycle, adapted from Guzik et al. (2020) [19].
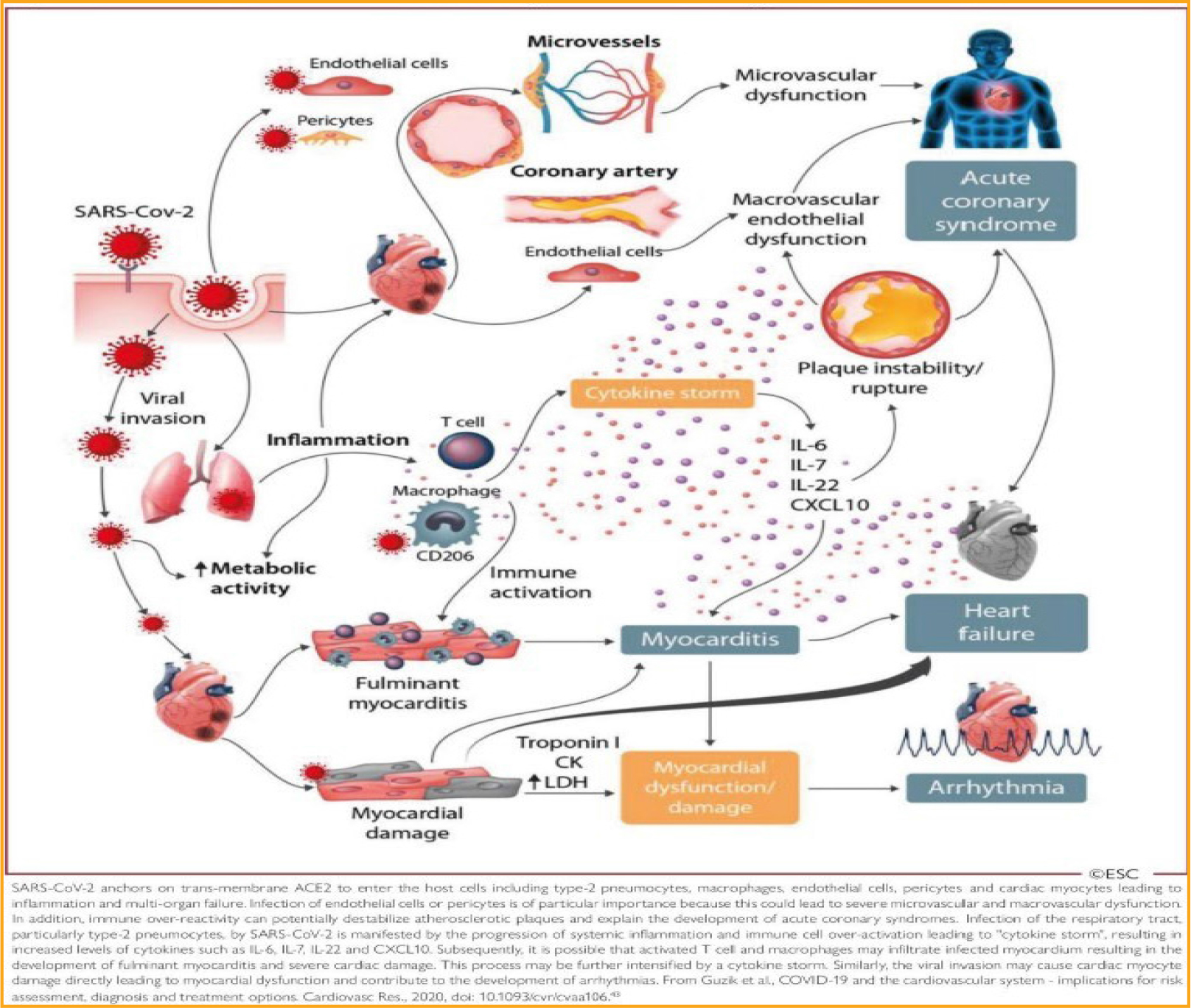
Figure 6: COVID-19 interaction with human pathophysiology, adapted from Guzik et al. (2020) [19].
Figure 7: Separation of mild and severe COVID-19 cases. Screenshot from VisiRule 7.021.
In a study of 416 COVID-19 patients, of whom 57 died of the illness, Shi et al. [40] found that cardiac injury was most common. Some research on COVID-19 infections has shown that interstitial inflammatory cells can infiltrate the myocardium, leading to congestive heart failure [31]. Indeed, COVID-19 infections are likely associated with infection-induced myocarditis and ischemia [41]. Moreover, in multivariable adjusted models, cardiac injury was significantly associated with mortality [24]. In a study by Guo et al. [39], elevated troponin T levels due to cardiac injury were associated with a significant rise in mortality. Severe COVID-19 infections can be associated with cardiac arrhythmias, at least in part because infection-related myocarditis and increased Troponin T levels result in damage to the heart’ muscle tissue [29]. These findings were fully integrated in the decision tree with binary classifications of overactive immune responses and heart damage, which is one of the decision-support strengths.
Important knowledge mechanisms were incorporated into the flow of questions in the decision tree to form the COVID-19 decision support (CCDS) tool. In fact, CCDS has 959 lines of code with 212 nodes (questions) and 48 end points in total comprising 18 long recovery, 13 deaths, 10 chronic disease onset with recovery, and 7 recovery. This coding amount covered the following main knowledge details:
- The pathobiology of COVID-19 binding to the host receptor angiotensin-converting enzyme 2 (ACE2), which mediates its entry into host cells, which is expressed in the lungs, heart, and blood vessels, is a key member of the renin angiotensin system (RAS), which plays an important role in the onset of cardiovascular diseases (CVD) [19];
- Cardiovascular disease associated with COVID-19 likely involves dysregulation of the RAS/ACE2 system by the infection, especially when comorbidities, such as hypertension [42], are present [43];
- CVD may be a primary phenomenon in COVID-19, but it may also be secondary to acute lung injury, which increases cardiac workload, a condition that is especially problematic in patients with congestive heart failure [26];
- A cytokine storm, resulting from a combination of T-cell activation and dysregulated release of interleukin (IL)-6, IL-17 and other cytokines, may contribute to CVD in COVID-19 cases [19]; and,
- Immune system activation may result in plaque instability, increasing the risk of acute coronary events such as stroke [26].
The host receptor through which COVID-19 enters cells to trigger infection is ACE2 (Figure 5). This process facilitates its entry into alveolar epithelial cells within the cytoplasm of potential host cells in the lung. The viral RNA then starts to replicate, followed by viral shedding, which likely plays a pathogenic role, resulting in severe cases in lung injury and respiratory failure [43]. In addition to the lungs, ACE2 is highly expressed in the human heart, blood vessels, and gastrointestinal tract [19]. These biological facts are well represented in the decision-support tool (Table 6).
A major strength of decision trees in the decision-support tool is their usefulness in the stratification of comorbidities. Comorbidities identified in COVID-19 studies include chronic cardiac disease, chronic respiratory disease, chronic renal disease (estimated glomerular filtration rate ≤30), mild to severe liver disease, dementia, chronic neurological conditions, connective tissue disease, Diabetes mellitus, and various malignancies [2, 31]. Clinician-defined obesity is also classed as a comorbidity owing to its probable association with adverse outcomes in patients with COVID-19 in New York City [31]. The decision tree can represent this stratification of comorbidities by increasing the depth from the trunk to the leaf endpoints.
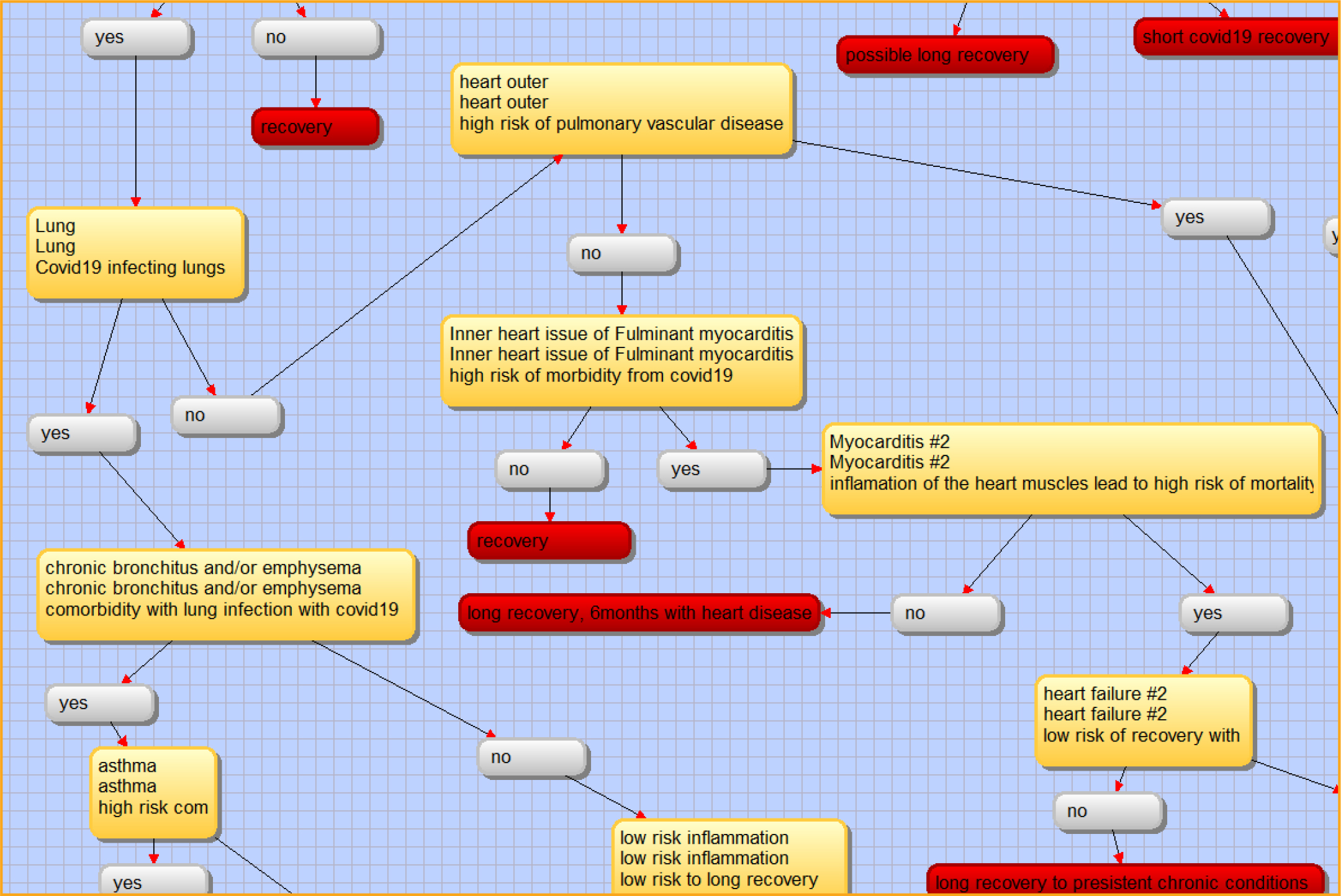
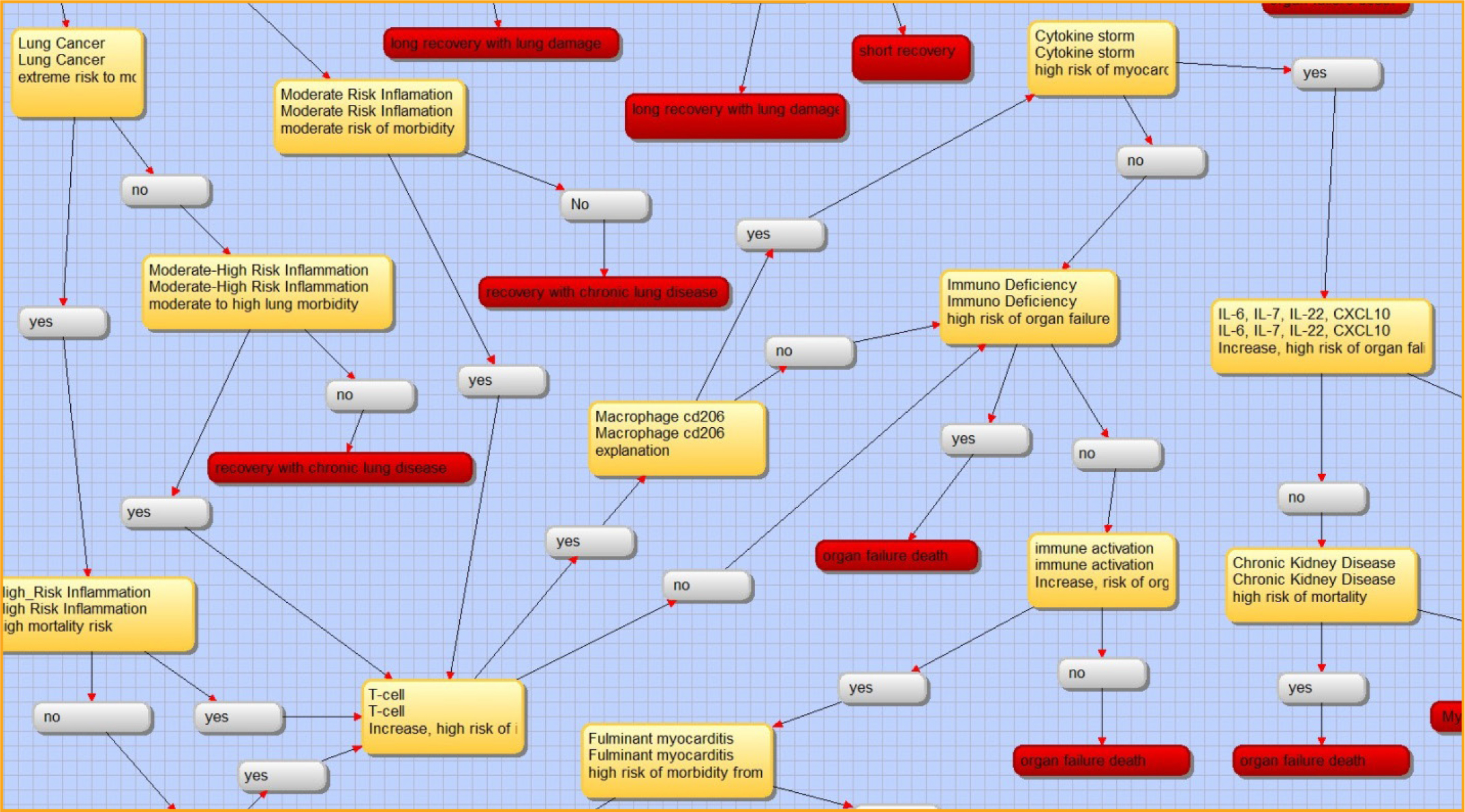
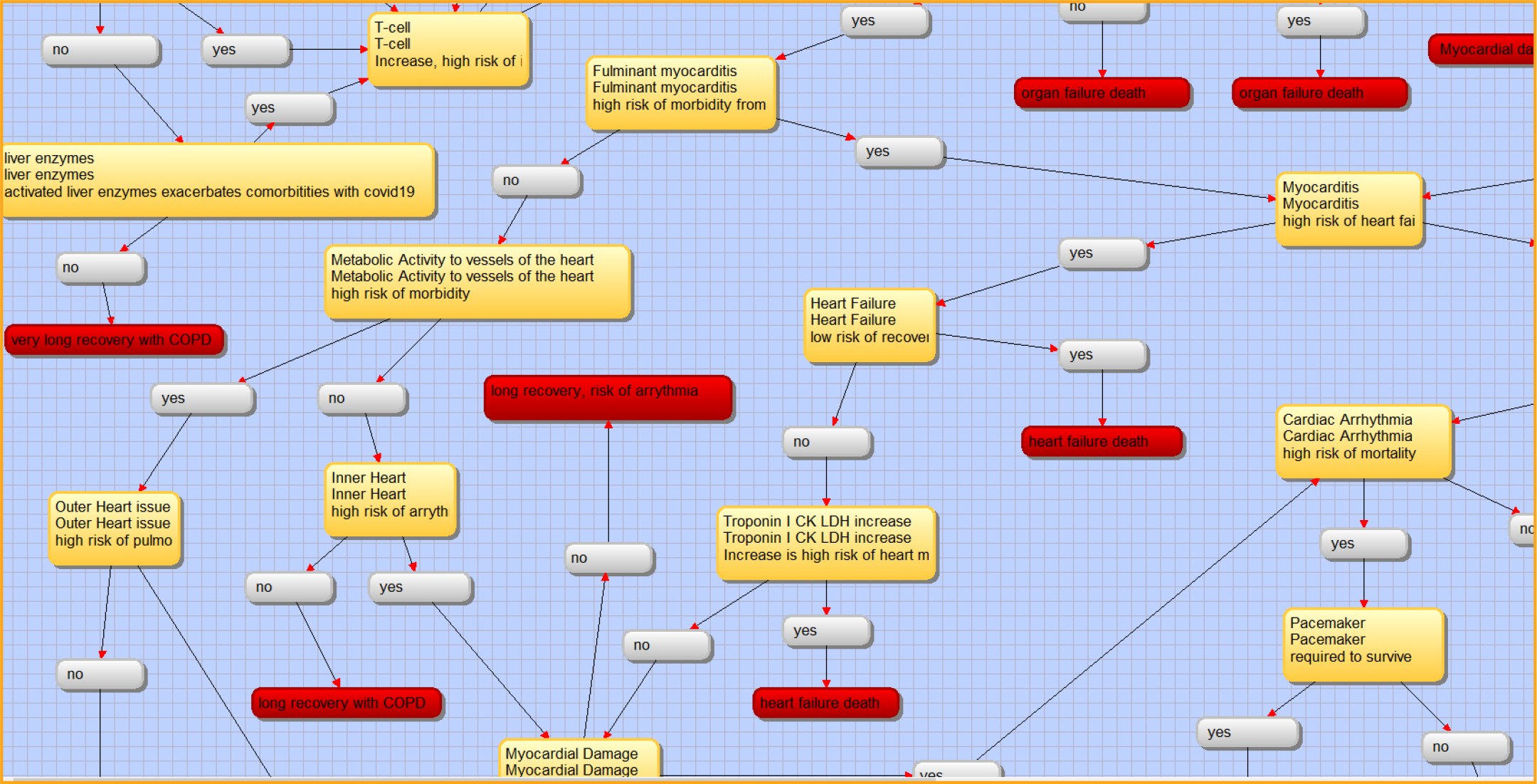
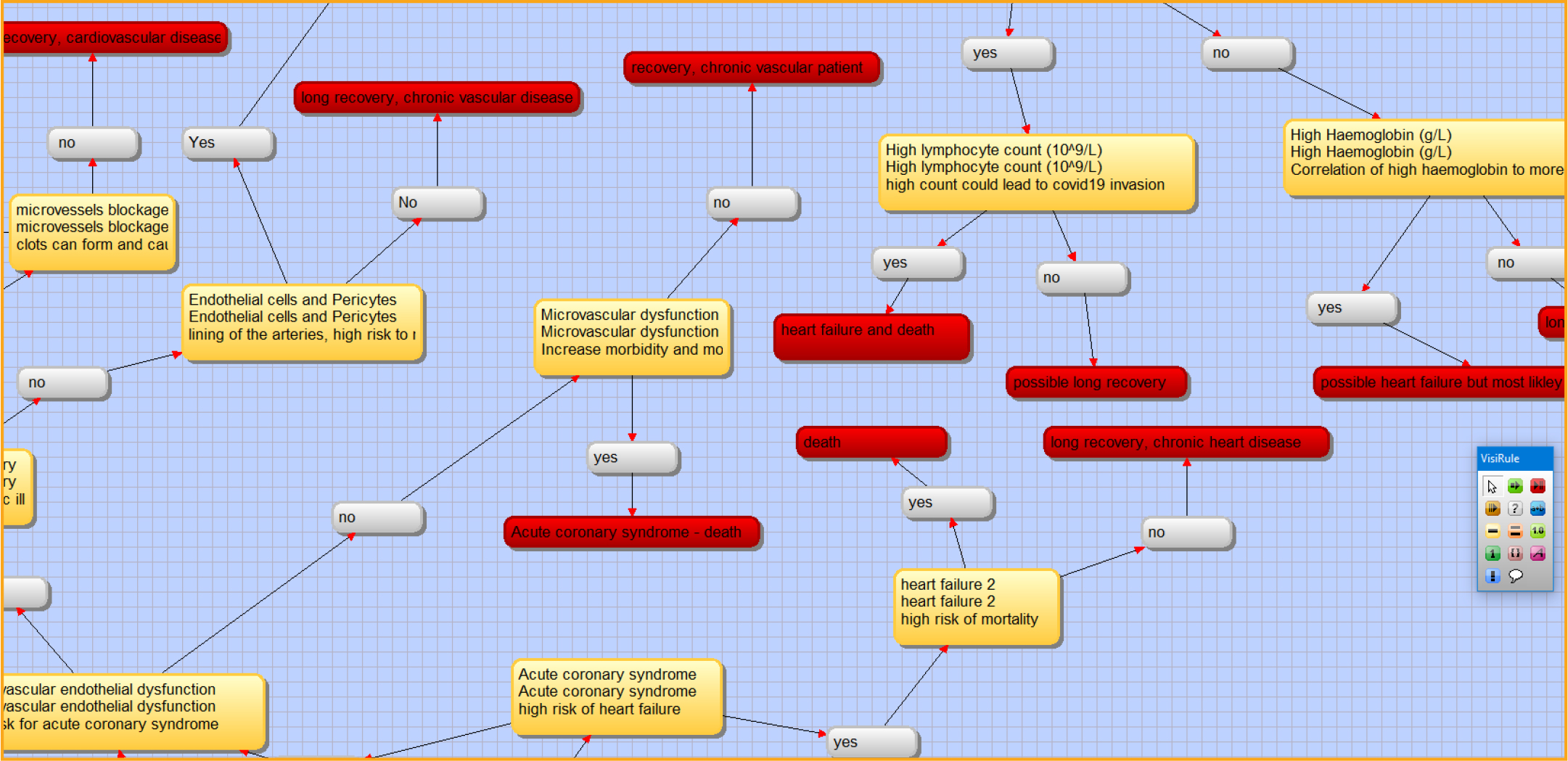
COVID-19 is primarily a respiratory disease, but many patients also develop cardiovascular disease, including hypertension [45, 46], acute cardiac injury, and myocarditis (Figure 6) [19]. The decision-support tool using a decision-tree framework can easily represent the transition from respiratory to cardiovascular diseases (Figure 8). Furthermore, the decision tree can improve decision support by modeling the patient’s varied immune responses, including inflammation, macro-vascular and microvascular dysfunctions, and multi-organ failure (Figure 6) [19]. These phenomena were well integrated in the clinical decision-support tool and important in binary classification of cases with high risk of mortality. Subsequently, it is possible that activated T cells and macrophages may infiltrate infected myocardium, resulting in the development of fulminant myocarditis and severe cardiac damage (Figure 9) [46–48]. This eventuality too is fully integrated in the decision-support tool, with several linkages in the decision tree between cytokine storms, arrhythmias, microvascular dysfunction, and acute coronary syndrome (Figures 10–11) [46]. Each biological response is represented by a node or leaf in the decision tree, with the stratification of health conditions correlating to specific levels of biological response.
Figure 8: Chronic lung conditions or inner heart. Screenshot from VisiRule 7.021.
Figure 9: Lung to Inflammation to T-cell and cytokine storm. Screenshot from VisiRule 7.021.
Figure 10: Troponin, Myocardial Damage, Cardiac Arrhythmia. Screenshot from VisiRule 7.021.
Figure 11: Microvascular dysfunction and acute coronary syndrome. Screenshot from VisiRule 7.021.
In a previous study, data mining models were developed for the prediction of COVID-19 patients’ recovery using an epidemiological dataset of COVID-19 patients in South Korea [28]. In that study, a decision tree, a support vector machine, naïve Bayes classifiers, logistic regression, a random forest, and a K-nearest neighbour algorithm were applied directly to the dataset, and a model was developed using the Python. The model predicted the age ranges of patients who are at high risk of dying from COVID-19, of those who are likely to recover, and of those who are likely to recover rapidly [28]. Additionally, literature indicates that COVID-19 positivity is associated with lymphopenia (that is, abnormally low levels of white cells in the blood), damage to liver and muscle tissue, and significantly increased levels of C-reactive protein (CRP) [49]. The results of the present study show that a model developed with a decision-tree algorithm can be most efficient in predicting the probability of recovery for COVID-19 patients.
Limitations
The use of clinical decision-support tools that incorporate decision trees is subject to an inherent limitation: the lack of circular reference, which is not possible in decision trees. In terms of best practice, the diagnosis of severe cases of COVID-19 with comorbidities requires circular references and feedback loops in a system-dynamic approach, especially with regard to pathophysiology. Furthermore, a list of frequent abnormalities in COVID-19 patients includes increased aspartate aminotransferase, decreased lymphocyte count, increased lactate dehydrogenase, increased CRP, increased white blood cell count, etc…[24, 50]. This suggests that future models of COVID-19 could incorporate some or all of these features; however, this shows a limitation of the decision-support tool. For example, a study highlighted the importance of being able to predict COVID-19 severity on the basis of blood samples, especially when these are analyzed using Gini-coefficients for recovery [28]. According to a recent announcement, blood tests used in diagnosing congestive heart failure can also help indicate severe cases of COVID-19 [51]. In my research, stratification of 6–7 leaves was carried out to include the relevant parameters for diagnosis and prognosis of COVID-19 [30]. However, the decision tree used in this decision-support tool did not set a threshold on the number of leaves per node and instead added knowledge structures of COVID-19 with human anatomy and biology.
Gender was not included as a variable in the decision tree. Patients are more likely to be men and to have more comorbidities such as hypertension and coronary heart disease [11, 40]. Men have a higher risk of cardiovascular complications than women do. Also, no geographical data were included, and no sensitivity analysis was applied, further limiting the usefulness of the study.
Another limitation is the application of inference probabilities of Bayes Theorem to the leaves of the decision tree were not interactive and static. There are many uncertainties in the health outcomes of patients with severe COVID-19; therefore, the application of Bayes probabilities is important. With the lack of success in automated construct of a decision tree from COVID-19 data, there was no method to automate a valid application of Bayes probabilities for risk of mortality versus recovery. The structure of decision trees allows for knowledge acquisition and application of inference probabilities but this cannot be validated at this time.
Moreover, if the decision tree is primarily ontology-based on its binary classification, then the probabilities of COVID-19 could become more accurate and plausible based on medical conditions such as co-morbidities. Kan, Saeidlou and Saadat (2019) showed a possible way to integrate a trained dataset using WEKA, MatLab then integrate the dataset with ontology via Protégé application to establish an ontology-based decision tree model [52]. This method could be applied to this COVID-19 decision tree and integrate ontology in its stratification, which would reduce the number of possible scenario at 63,360 to a much lower number but the scenarios would have incorporated ontology rules like age and body system with co-morbidities more succinct. In turn, a dataset could be simulated and ratified towards the type of data that needs to be collected to form a similar decision tree for accurate binary classification of stratified severe COVID-19 cases.
Finally, the use of decision trees makes it difficult to control for certain biases. Decision trees use induction-to-deduction algorithms that range from traditional heuristic-based techniques to more recent hybrid data-to-tree approaches. These algorithms are essential in constructing a sequence of questions that flow from one to the next. For this reason, the basic features of the decision-support tool mitigate against the quantification of any inherent biases. For example, the stratification could be biased by the decision to design a sequence from chronic lung conditions to the heart instead of to gastrointestinal illnesses. This places an emphasis on knowledge of the cardiovascular system while underlying conditions affecting other body systems.
References
- Wölfel R, Corman VM, Guggemos W, et al. (2020) Virological assessment of hospitalized patients with COVID-19. Nature 581: 465–469. [View]
- Yang X, Yu Y, Xu J, et al. (2020) Clinical course and outcomes of critically ill patients with SARS- CoV-2 pneumonia in Wuhan, China: a single-centered, retrospective, observational study. Lancet Respir Med 8: 475–81. [View]
- Dong E, Du H, Gardner L (2020) An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect Dis S1473- 3099(20)30120–1. [View]
- Johns Hopkins (2020) Coronavirus Resource Center and COVID-19 map. [View]
- Government of Canada (2020) Coronavirus disease (COVID-19). [View]
- Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D (2020) Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 181: 281–292 e6. [View]
- Yan R, Zhang Y, Li Y, Xia L, Guo Y, Zhou Q (2020) Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367: 1444–1448. [View]
- van Doremalen N, Bushmaker T, Morris DH, Holbrook MG, Gamble A, (2020) Aerosol and Surface Stability of SARS-CoV-2 as Compared with SARS-CoV-1. New England J Medicine 382: 1564–1567. [View]
- Ai T, Yang Z, Hou H, Zhan C, Chen C, et al. (2020) Correlation of Chest CT and RT-PCR Testing in Coronavirus Disease 2019 (COVID-19) in China: A Report of 1014 Cases. Radiology 2020: 200642. [View]
- Wynants L, Calster BV, Collins GS, Riley RD, Heinze G, et al. (2020) Prediction models for diagnosis and prognosis of COVID-19: systematic review and critical appraisal. BMJ 369: m1328. [View]
- Zhou F, Yu T, Du R, Fan G, Liu Y, et al. (2020) Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 395: 1054–1062. [View]
- Roozenbeck J, Schneider CR, Dryhurst S, Kerr J, Freeman ALJ, et al. (2020) Susceptibility to misinformation about COVID-19 around the world. R Soc Open Sci 7: 201199. [View]
- Driggin E, Madhavan MV, Bikdeli B, Chuich T, Laracy J, Biondi-Zoccai G, et al. (2020) Cardiovascular Considerations for Patients, Health Care Workers, and Health Systems During the COVID-19 Pandemic. J Am Coll Cardiol 75: 2352–2371. [View]
- Moutounet-Cartan PGB (2020) Deep convolutional neural networks to diagnose COVID-19 and other pneumonia diseases from posteroanterior chest x-rays. ArXiv e-prints [Preprint] Cornell University. https://arxiv.org/abs/2005.00845 Accessed November 1, 2020.
- Gao Y, Cai GY, Fang W, Li WY, wang ST, Chen L, et al. (2020) Machine learning based early warning system enables accurate mortality risk prediction for COVID-19. Nature Communications 11: 5033. [View]
- Diaz-Quijano FA, Silva JMN, Ganem F, et al. (2020) A model to predict SARSCoV-2 infection based on the first three-month surveillance data in Brazil. Tropical Medicine and International Health 25: 1385–1394. [View]
- Shi H, Han X, Jiang N, Cao Y, Alwalid O, Gu J, Fan Y, Zheng C (2020) Radiological findings from 81 patients with COVID-19 pneumonia in Wuhan, China: a descriptive study. Lancet Infect Dis 20: 425–434. [View]
- Brinati D, Campagner A, Ferrari D, Locatelli M, Banfi G, Cabitza F (2020) Detection of COVID-19 Infection from Routine Blood Exams with Machine Learning: A Feasibility Study. Journal of Medical Systems 44 (135). [View]
- Guzik TJ, Mohiddin SA, Dimarco A, Patel V, Savvatis K, et al. (2020) COVID-19 and the cardiovascular system: implications for risk assessment, diagnosis, and treatment options. Cardiovasc Res 116: 1666–1687. [View]
- Zeng J, Huang J, Pan L (2020) How to balance acute myocardial infarction and COVID-19: the protocols from Sichuan Provincial People’s Hospital. Intensive Care Med 46: 1111–1113. [View]
- The RECOVERY Collaborative Group (2020) Dexamethasone in hospitalized patients with Covid- 19 — preliminary report. N Engl J Med. NEJMoa2021436. [View]
- McRae MP, Dapkins IP, Sharif I, Anderman J, Fenyo D, et al. (2020) Managing COVID-19 with a clinical decision support tool in a community health network: algorithm development and validation. J Medical Internet Research 22: e22033: 1–13. [View]
- Xiong TY, Redwood S, Prendergast B, Chen M (2020) Coronaviruses and the cardiovascular system: acute and long-term implications. Eur Heart J 41: 1798–1800. [View]
- Knight SR, Ho A, Pius R, Buchan I, Carson G, et al. (2020) Risk stratification of patients admitted to hospital with COVID-19 using the ISARIC WHO Clinical Characterisation Protocol: development and validation of the 4C Mortality Score. BMJ 370: m3339. [View]
- Oyelade O, Ezugwu AE (2020) A case-based reasoning framework for early detection and diagnosis of novel coronavirus. Informatics in Medicine 20: 100395. [View]
- Madjid M, Safavi-Naeini P, Solomon SD, Vardeny O (2020) Potential Effects of Coronaviruses on the Cardiovascular System: A Review. JAMA Cardiol 5: 831–840. [View]
- Wu Y (2020) Compensation of ACE2 Function for Possible Clinical Management of 2019-nCoV- Induced Acute Lung Injury. Virol Sin 35: 256–258. [View]
- Muhammad LJ, Islam M, Usman SS, Ayon SI (2020) Predictive Data Mining Models for Novel Coronavirus (COVID?19) Infected Patients’ Recovery. SN Computer Science 1: 206. [View]
- Flores F, Walter J, Wussler D, Kozhuharov N, Nowak A, Dinort J, Badertscher P, et al. (2019) Direct comparison of high-sensitivity cardiac troponin t and i for prediction of mortality in patients with pneumonia. J Clin Chem Lab Med 2: 1000131. [View]
- Azar AT, El-Metwally E (2013) Decision tree classifiers for automated medical diagnosis. Neural Computer and Application 23: 2387–2403. [View]
- Richardson S, Hirsch JS, Narasimhan M, Crawford JM, McGinn T, Davidson KW, et al. (2020) Presenting Characteristics, Comorbidities, and Outcomes Among 5700 Patients Hospitalized With COVID-19 in the New York City Area. JAMA 323: 2052–2059. [View]
- VisiRule (2020) VisiRule Expert System Software – Medical Expert System Demos (COVID-19 by Dillon Chrimes). Accessed October 30, 2020. [View]
- Wiguna W, Riana D (2020) Diagnosis of coronavirus disease 2019 (COVID-19) surveillance using c4.5 algorithm. Journal PILARNusa Mandiri 16: 71–81. [View]
- Google Cloud (2020) Dialogflow. [View]
- Zenchat (2020) Chat + Tasks = Magic. [View]
- Espinoza J, Crown K, Kulkarni O (2020) A guide to chatbots for COVID-19 screening at pediatric health care facilities. JMIR Public Health Surveillance 6: e18808. [View]
- Albu A (2017) From logical inference to decision trees in medical diagnosis. The 6th IEEE International Conference on E-Health and Bioengineering - EHB 2017. Grigore T. Popa University of Medicine and Pharmacy, Sinaia, Romania, June 22–24, 2017. 65–68. [View]
- Podgorelec V, Kokol P, Stiglic B, Rozman I (2002) Decision trees: an overview and their use in medicine. J Med Syst 26: 445–63. [View]
- Guo T, Fan Y, Chen M, Wu X, Zhang L, et al. (2020). Cardiovascular Implications of Fatal Outcomes of Patients With Coronavirus Disease 2019 (COVID-19). JAMA Cardiol 5: 811–818. [View]
- Shi S, Qin M, Shen B, Cai Y, Liu T, et al. (2020) Association of Cardiac Injury With Mortality in Hospitalized Patients With COVID-19 in Wuhan, China. JAMA Cardiol 5: 802–810. [View]
- Giannitsis E, Mair J, Christersson C, Siegbahn A, Huber K, et al. (2017) Biomarker Study Group of the European Society of Cardiology Acute Cardiovascular Care A. How to use D-dimer in acute cardiovascular care. Eur Heart J Acute Cardiovasc Care 6: 69–80. [View]
- Fang L, Karakiulakis G, Roth M (2020) Are patients with hypertension and diabetes mellitus at increased risk for COVID-19 infection? Lancet Respir Med 8: e21. [View]
- Santos RAS, Sampaio WO, Alzamora AC, Motta-Santos D, Alenina N, et al. (2018) The ACE2/Angiotensin-(1–7)/MAS Axis of the Renin- Angiotensin System: Focus on Angiotensin-(1–7). Physiol Rev 98: 505–553. [View]
- Drummond GR, Vinh A, Guzik TJ, Sobey CG (2019) Immune mechanisms of hypertension. Nat Rev Immunol 19: 517–532. [View]
- Williams B, Zhang Y (2020) Hypertension, renin-angiotensin-aldosterone system inhibition, and COVID-19. Lancet 395: 1671–1673. [View]
- Roffi M, Patrono C, Collet JP, Mueller C, Valgimigli M, Andreotti F, Bax JJ, et al. (2016) 2015 ESC Guidelines for the management of acute coronary syndromes in patients presenting without persistent ST-segment elevation: Task Force for the Management of Acute Coronary Syndromes in Patients Presenting without Persistent ST-Segment Elevation of the European Society of Cardiology (ESC). Eur Heart J 37: 267–315. [View]
- Li Z, Guo X, Hao W, Wu Y, Ji Y, Zhao Y, et al. (2003) The relationship between serum interleukins and T-lymphocyte subsets in patients with severe acute respiratory syndrome. Chinese Medical Journal 116: 981–4. [View]
- Hu H, Ma F, Wei X, Fang Y (2020) Coronavirus fulminant myocarditis saved with glucocorticoid and human immunoglobulin. Eur Heart J 42: 206. [View]
- Siedlinski M, Jozefczuk E, Xu X, Teumer A, Evangelou E, et al. (2020) White Blood Cells and Blood Pressure: A Mendelian Randomization Study. Circulation 141: 1307–1317. [View]
- Wu C, Chen X, Cai Y, Xia J, Zhou X, Xu S, et al. (2020) Risk Factors Associated With Acute Respiratory Distress Syndrome and Death in Patients With Coronavirus Disease 2019 Pneumonia in Wuhan, China. JAMA Intern Med. [View]
- Tan L, Wang Q, Zhang D, Ding J, Huang Q, et al. (2020) Lymphpenia predicts disease severity of COVID-19: a descriptive and predictive study. Signal Transduction and Targeted Therapy 5: 1–3. [View]
- Khan ZMA, Saeidlou S, Saadat M (2019) Ontology-based decision tree model for prediction in a manufacturing network. Production & Manufacturing Research 7: 335–349. [View]
Article Type
Research Article
Publication history
Received: June 17, 2022
Accepted: June 28, 2022
Published: June 28, 2022
Citation:
Chrimes D, Zamani H, Spenser C, Westwood A (2022) Decision-Support Expert System to Assess Severe COVID-19. Clar J Infect Dis Ther 03(01): 207–231.
Dillon Chrimes1*, Hamid Zamani2, Clive Spenser3 and Alan Westwood3
1School of Health Information Science, Human and Social Development, University of Victoria, Victoria BC Canada
2Vancouver Island Health Authority, Corporate Business Solutions, Victoria BC Canada
3VisiRule, LPA Ltd, UK
*Corresponding author
Dillon Chrimes, PhD,
School of Health Information Science,
Department of Human and Social Development,
University of Victoria,
Canada;
Tel: 250-857-2404;